Disclaimer: This site is a part of curriculum project. The information should be used for educational purpose only.
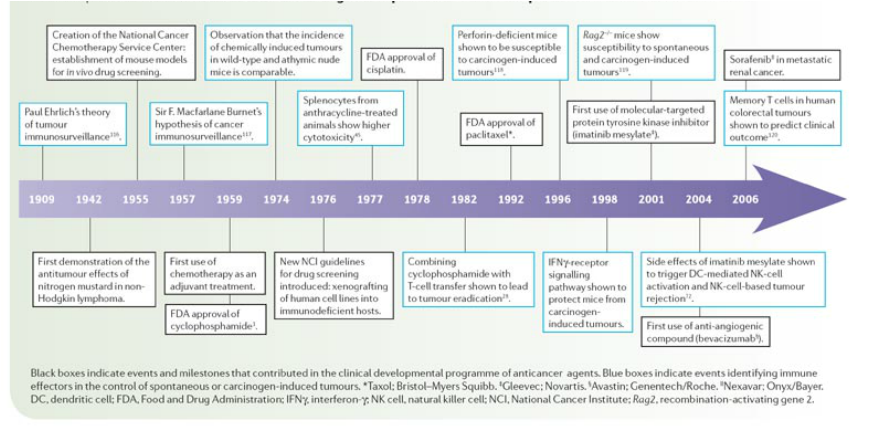 fig.11 Evolution of Anticancer Drug Development
fig.11 Evolution of Anticancer Drug Development
EVOLUTION OF ANTICANCER DRUG DISCOVERY
The approach to the discovery of new anticancer drugs has recently evolved from a reliance on empiric cell-based screening for antiproliferative effects to a more mechanistically based approach that targets the specific molecular lesions thought to be responsible for the development and maintenance of the malignant phenotype in various forms of cancer. The ultimate goal of the development of molecularly targeted drugs is to improve the efficacy and selectivity of cancer treatment by exploiting the differences between cancer cells and normal cells. The success of recently developed molecularly targeted agents, such as tretinoin (all-trans-retinoic acid) for acute promyelocytic leukemia and imatinib (STI-571) for chronic myelogenous leukemia (CML) and gastrointestinal stromal tumors , provides early clinical validation for the molecularly targeted approach to drug discovery. Most of the commonly used cytotoxic anticancer drugs were discovered through random high-throughput screening of synthetic compounds and natural products in cell-based cytotoxicity assays. Despite the number and chemical diversity of these agents, the mechanisms of action are limited , and most compounds are DNA-damaging agents with a low therapeutic index. With this screening approach, mechanism of action is not a primary determinant in selecting agents for further development, and, as a result, none of the current drugs directly targets the molecular lesions responsible for malignant transformation. The initial National Cancer Institute (NCI) high-throughput screen used the highly chemosensitive P388 leukemia cell line, but this screen failed to identify drugs that were active against the common adult solid tumors. In the mid-1980s, the NCI implemented a new in vitro disease-oriented screen consisting of 60 human tumor cell lines representing nine common forms of cancer. It remains to be determined whether selective activity in vitro against cell lines representing a particular histologic form of cancer will be predictive for antitumor activity in vivo . As the molecular bases of specific forms of cancer are elucidated, a variety of new potential therapeutic targets are being identified. In the most common forms of cancers, the accumulation of mutations in multiple genes is required for tumorigenesis . These mutations, such as the ras-activating mutations and mutations that inactivate p53, allow cancer cells to circumvent intrinsic and extrinsic controls that tightly regulate the cell cycle and cell division and apoptosis in normal cells. After identification of the genetic alterations in cancer cells, the critical transforming mutations must be discerned from genetic alterations that are the result of the inherent genetic instability in cancer cells but that do not play a direct role in tumorigenesis . This target validation is performed in preclinical cancer models. Identification of valid molecular targets has led to rational target-based drug discovery at the protein level, as illustrated by the development of imatinib, which was discovered by screening compound libraries for inhibitors of the protein kinase activity in vitro . Many of the proteins involved in cell cycle regulation, signal transduction, and the regulation of apoptosis are enzymes or receptors and are, therefore, potentially amenable to inhibition by small molecules Finally, most molecular targets for new cancer treatments interact with other proteins within pathways or networks in the cell, and the pharmacologic effect resulting from the inhibition of a specific target may be influenced by the expression or relative levels of these interacting proteins. Therefore, target-based screening assays may not be predictive of drug effect within the context of the whole cell . As demonstrated by Dunstan et al. cell-based screening assays will continue to play an important role in drug discovery as we move into the moleculartargeting era. Their three-stage cell-based procedure using yeast cells that can be genetically manipulated is adaptable to highthroughput screening to identify agents that have a selective effect against cells with specific mutations (gene deletions). Unlike target-based assays, this cell-based assay is not a mechanistic screen, but determining the mechanism of action of selectively toxic agents from this screen may identify new molecular targets. (Journal of National Cancer Institue, Vol.94, Jan.16 2002. Frank Balis,2002)
REFERENCE:- Journal of the National Cancer Institute, Vol. 94, No. 2, January 16, 2002 -
Evolution of Anticancer Drug Discovery by Frank M. Balis
The approach to the discovery of new anticancer drugs has recently evolved from a reliance on empiric cell-based screening for antiproliferative effects to a more mechanistically based approach that targets the specific molecular lesions thought to be responsible for the development and maintenance of the malignant phenotype in various forms of cancer. The ultimate goal of the development of molecularly targeted drugs is to improve the efficacy and selectivity of cancer treatment by exploiting the differences between cancer cells and normal cells. The success of recently developed molecularly targeted agents, such as tretinoin (all-trans-retinoic acid) for acute promyelocytic leukemia and imatinib (STI-571) for chronic myelogenous leukemia (CML) and gastrointestinal stromal tumors , provides early clinical validation for the molecularly targeted approach to drug discovery. Most of the commonly used cytotoxic anticancer drugs were discovered through random high-throughput screening of synthetic compounds and natural products in cell-based cytotoxicity assays. Despite the number and chemical diversity of these agents, the mechanisms of action are limited , and most compounds are DNA-damaging agents with a low therapeutic index. With this screening approach, mechanism of action is not a primary determinant in selecting agents for further development, and, as a result, none of the current drugs directly targets the molecular lesions responsible for malignant transformation. The initial National Cancer Institute (NCI) high-throughput screen used the highly chemosensitive P388 leukemia cell line, but this screen failed to identify drugs that were active against the common adult solid tumors. In the mid-1980s, the NCI implemented a new in vitro disease-oriented screen consisting of 60 human tumor cell lines representing nine common forms of cancer. It remains to be determined whether selective activity in vitro against cell lines representing a particular histologic form of cancer will be predictive for antitumor activity in vivo . As the molecular bases of specific forms of cancer are elucidated, a variety of new potential therapeutic targets are being identified. In the most common forms of cancers, the accumulation of mutations in multiple genes is required for tumorigenesis . These mutations, such as the ras-activating mutations and mutations that inactivate p53, allow cancer cells to circumvent intrinsic and extrinsic controls that tightly regulate the cell cycle and cell division and apoptosis in normal cells. After identification of the genetic alterations in cancer cells, the critical transforming mutations must be discerned from genetic alterations that are the result of the inherent genetic instability in cancer cells but that do not play a direct role in tumorigenesis . This target validation is performed in preclinical cancer models. Identification of valid molecular targets has led to rational target-based drug discovery at the protein level, as illustrated by the development of imatinib, which was discovered by screening compound libraries for inhibitors of the protein kinase activity in vitro . Many of the proteins involved in cell cycle regulation, signal transduction, and the regulation of apoptosis are enzymes or receptors and are, therefore, potentially amenable to inhibition by small molecules Finally, most molecular targets for new cancer treatments interact with other proteins within pathways or networks in the cell, and the pharmacologic effect resulting from the inhibition of a specific target may be influenced by the expression or relative levels of these interacting proteins. Therefore, target-based screening assays may not be predictive of drug effect within the context of the whole cell . As demonstrated by Dunstan et al. cell-based screening assays will continue to play an important role in drug discovery as we move into the moleculartargeting era. Their three-stage cell-based procedure using yeast cells that can be genetically manipulated is adaptable to highthroughput screening to identify agents that have a selective effect against cells with specific mutations (gene deletions). Unlike target-based assays, this cell-based assay is not a mechanistic screen, but determining the mechanism of action of selectively toxic agents from this screen may identify new molecular targets. (Journal of National Cancer Institue, Vol.94, Jan.16 2002. Frank Balis,2002)
REFERENCE:- Journal of the National Cancer Institute, Vol. 94, No. 2, January 16, 2002 -
Evolution of Anticancer Drug Discovery by Frank M. Balis
